Duneau Lab
Evolutionary Immunology & Host-Pathogen Interactions
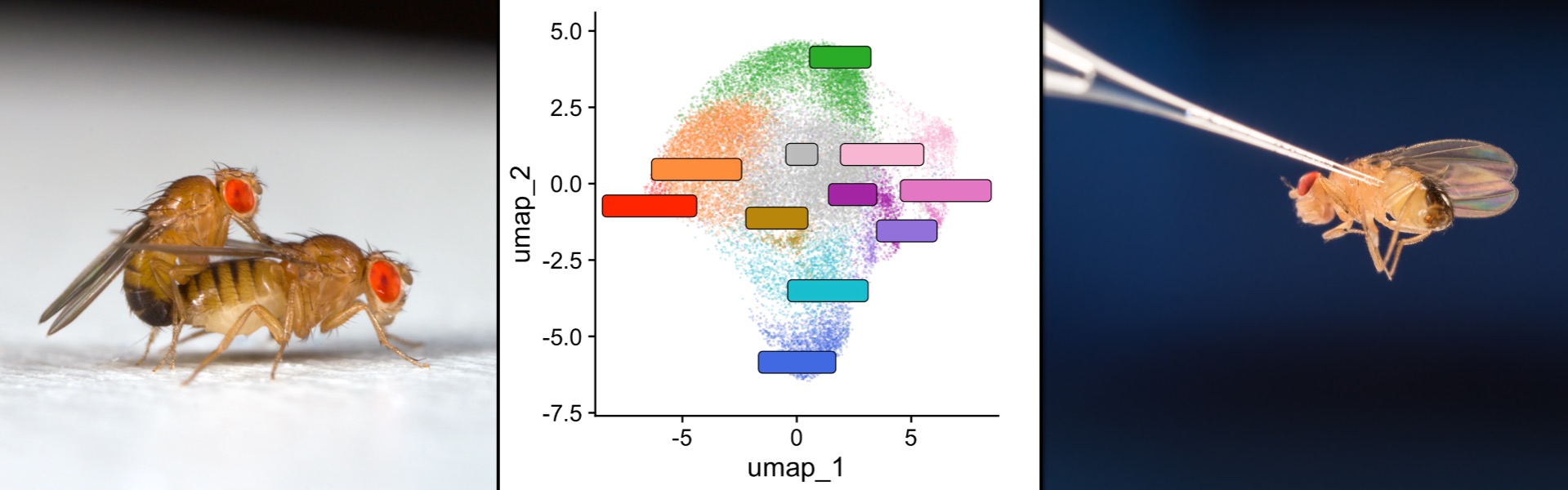
Our lab seeks to understand why individuals — and in particular males and females — respond differently to infection, and how this variation fuels the evolution of infectious diseases. We combine functional genetics, genomics, and mathematical modelling to study host-parasite interactions and how the environment shapes them.
Projects

Sexual dimorphism of diseases
Most infectious diseases have a sexually dimorphic outcome. We investigate the reasons for this not only with a host-centered view but also by considering that pathogens adapt to the most commonly encountered sex.

Host manipulation by parasites
One of the parasite strategy to increase its transmission is to manipulates its host. I started science by studying such strategy.
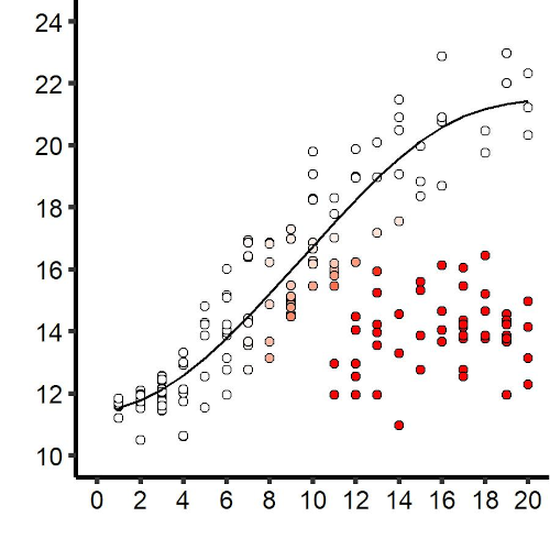
Within-host dynamics and disease outcomes
Infection outcome depends on the success of the parasite within its host. With theoretical and empirical approaches, we study what influences pathogen dynamics and its implication on symptoms.
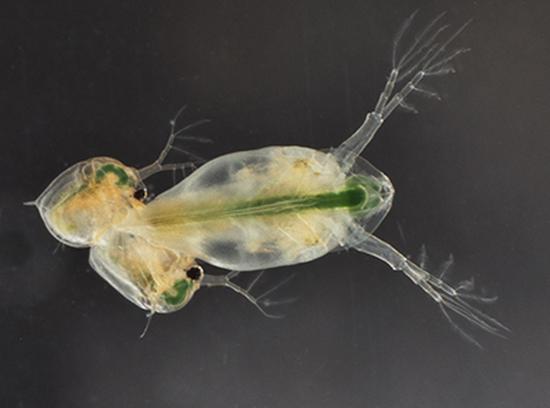

Sexual selection and conflicts
In the interest of better understanding the differences between sexes, we have studied sexual selection and conflict. We did so in two ways:
I- Parthenogenetic species produce mainly daughters without males. We study selection during the rare events of sexual reproduction in Daphnia.
II- Seminal fluid proteins are generally considered male tools to manipulate females. We investigated the growing hypothesis that seminal fluid proteins are important signals for females to adjust their egg-laying and mating decisions, and thus help reduce sexual conflict.
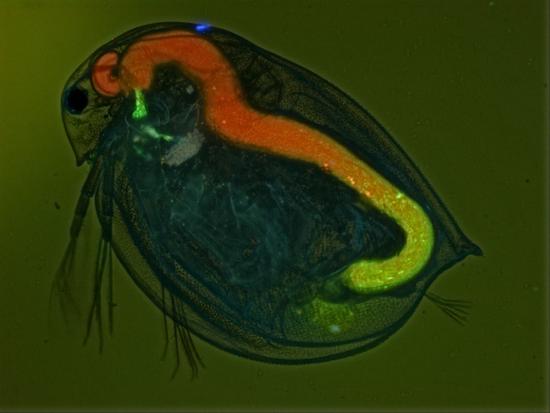
Mechanism host-parasite coevolution
Parasites need to go through several infection steps to be successfull. We study how the evolution at each step shapes host-parasite coevolution.

Phenotypic plasticity as generator of phenotypic variation
Genetic variation is the raw material for evolution. We pertain to study how phenotypic plasticity generates phenotypic heterogeneity, its genetic basis, and how it can explain variation in host-parasite interactions.
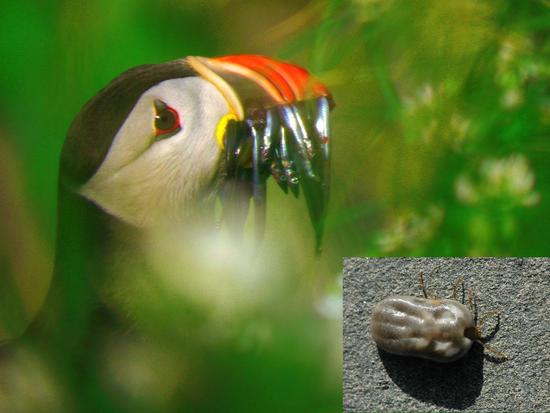

Lyme disease in seabirds
Lyme disease is a major threat in many countries. The bacteria responsible for it is mainly transmitted by ticks feeding on mammals. Long ago, I studied the role of seabirds as reservoir.
Featured publications
A within-host infection model to explore tolerance and resistance
Developed a within-host infection model integrating theory and experiments to distinguish between tolerance and resistance mechanisms. Provided a practical experimental framework for explaining individual differences in susceptibility to infection, a methodological foundation applied in subsequent work.
Gut cancer increases the risk of Drosophila being preyed upon by hunting spiders.
Diseased individuals are more preyed upon than healthy individuals. Although this is intuitive, this conventional wisdom has been subjected more to speculation than to empirical study. To test the idea, we genetically induced colon cancer in Drosophila and found that cancerous flies are more frequently predated than healthy ones.
Stochastic variation in the initial phase of bacterial infection predicts the probability of survival in Drosophila melanogaster.
A central problem with biomedicine is to understand why two individuals exposed to seemingly identical infections may have radically different clinical outcomes. Using the Drosophila melanogaster model, we analyse in depth, both through functional genetics and mathematical modelling, the main determinants that underlie the stochastic outcome of infection.
Host sexual dimorphism and parasite adaptation.
In this “essay” we propose for the first time the idea that the sexual dimorphism of diseases may be the result of the specific adaptation of parasites to the sex of their host. Similarly, as organisms adapt to the environment to which they are most frequently exposed, parasites can adapt to the sex they encounter most frequently (e.g., either because males and females are exposed differently, or because one sex is more easily infected than another due to immune differences). As a result, parasites behave differently depending on the sex they infect.
Publications
Five most recents
Sex differences in Drosophila intestinal metabolism contribute to sexually dimorphic infection outcome and alter gut pathogen virulence
We studied the sexual dimorphism of intestinal infections of D. melanogaster. We showed that males exhibit increased a key antioxidant defence system. It allows them to withstand oxidative stress-induced defecation blockage and clear the pathogen from the intestine, resulting in survival. We also showed that the bacteria showed increased expression of several virulence factors in female gut, indicating a change in pathogen behaviour depending on the host they infect (a theory I have been developing since Duneau & Ebert PLoS Biol. 2012). Contribution: I trained the first author to perform the analyses, and I analysed the GWAS.
Seminal fluid proteins as regulation factors for optimizing reproduction: a modeling approach.
Developed a biologically informed theoretical model showing how seminal fluid proteins synchronize sperm and egg release, reducing unfertilized egg production. Identified SP exhaustion as the key synchronization signal and demonstrated that sexual conflict over SP-mediated regulation of female physiology should be limited, though conflict over optimal remating rate remains possible.
Wound-induced eyespots on butterfly wings at the intersection of immune response and pigmentation development
Showed that immune activation modulates wound-induced ectopic eyespot formation in butterfly wings, revealing crosstalk between immune function and pigmentation development.
A within-host infection model to explore tolerance and resistance
Developed a within-host infection model integrating theory and experiments to distinguish between tolerance and resistance mechanisms. Provided a practical experimental framework for explaining individual differences in susceptibility to infection, a methodological foundation applied in subsequent work.
Genetic architecture of plasticity for pigmentation components in Drosophila melanogaster.
Applied genome-wide association mapping in Drosophila melanogaster to dissect the genetic basis of temperature-dependent pigmentation plasticity. Found that different body parts rely on largely independent genetic architectures, challenging the assumption of a common genetic basis for pigmentation plasticity. Contribution: I supervised the student for the genomic analysis and for the validation of allele candidate with functional genetics.
Contact
- dduneau@ciencias.ulisboa.pt
- University of Lisbon (FCUL)
1749-016 Lisboa - C2 Building
- DDuneau
